
The Statistical Analysis of Compositional Data: 9781930665781: Medicine & Health Science Books @ Amazon.com
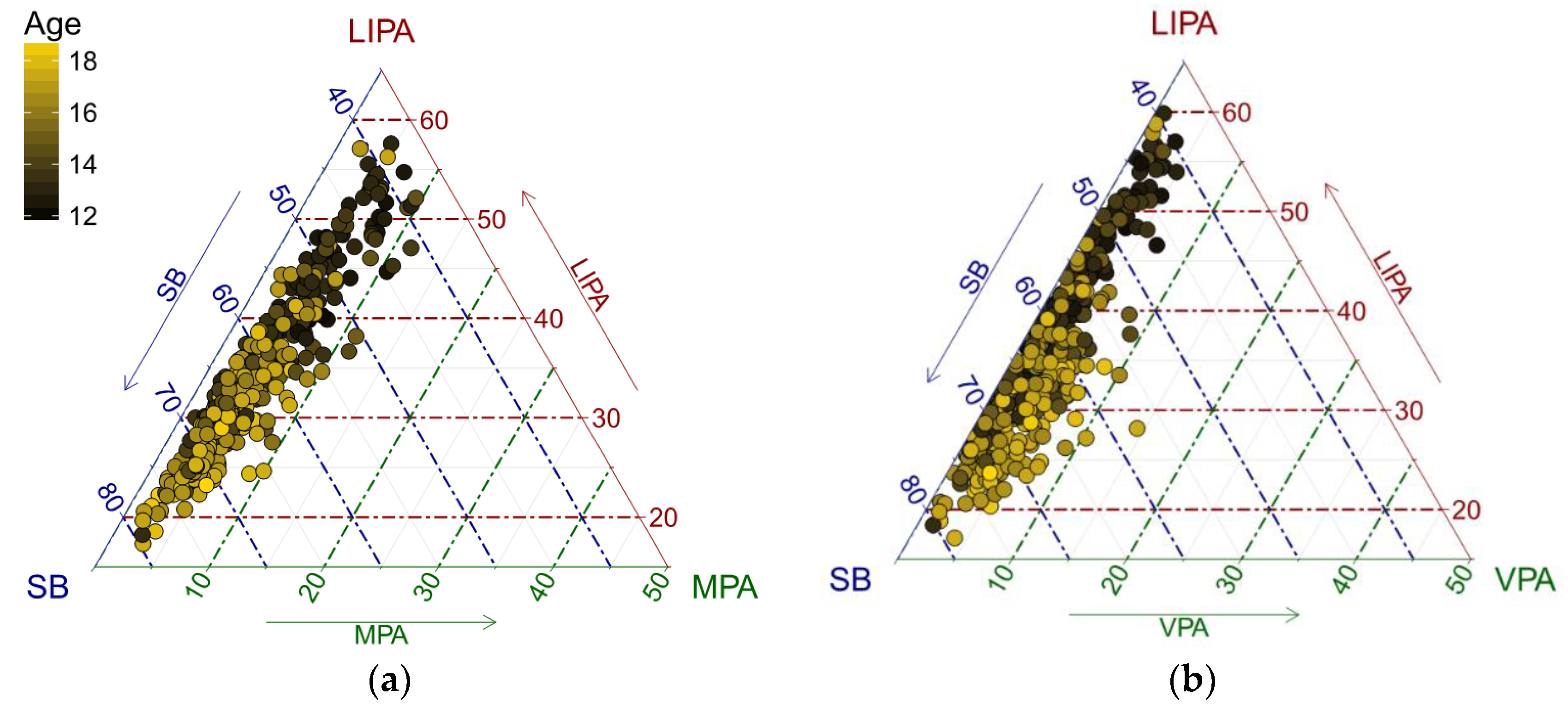
IJERPH | Free Full-Text | Robust Compositional Analysis of Physical Activity and Sedentary Behaviour Data
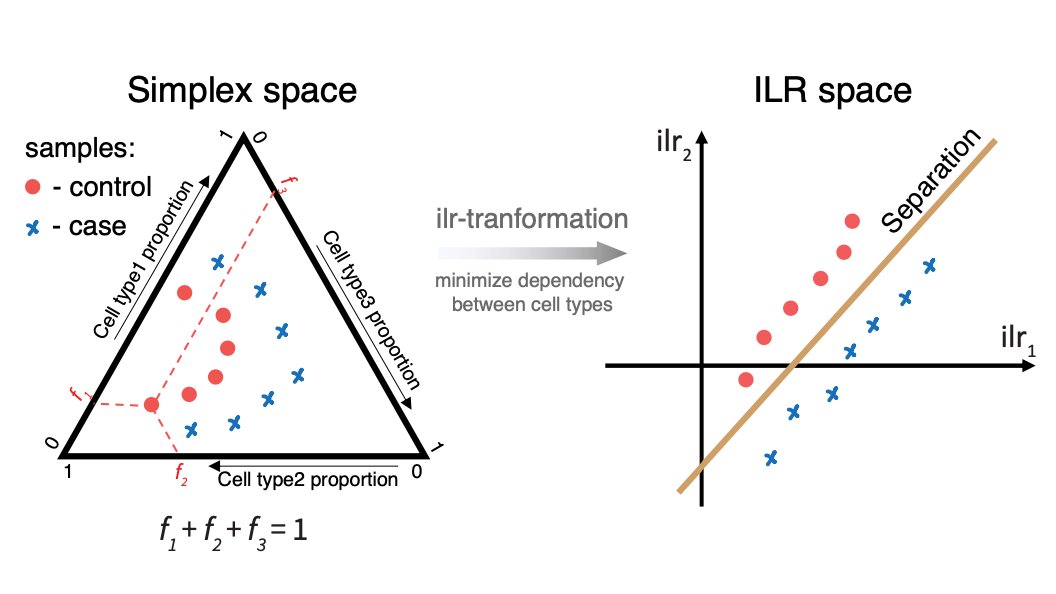
PeterK on Twitter: "For cluster-based compositional analysis we built on classic Compositional Data Analysis (CoDA) techniques, testing for statistical contribution of different cell types to a surface in simplex space optimally separating
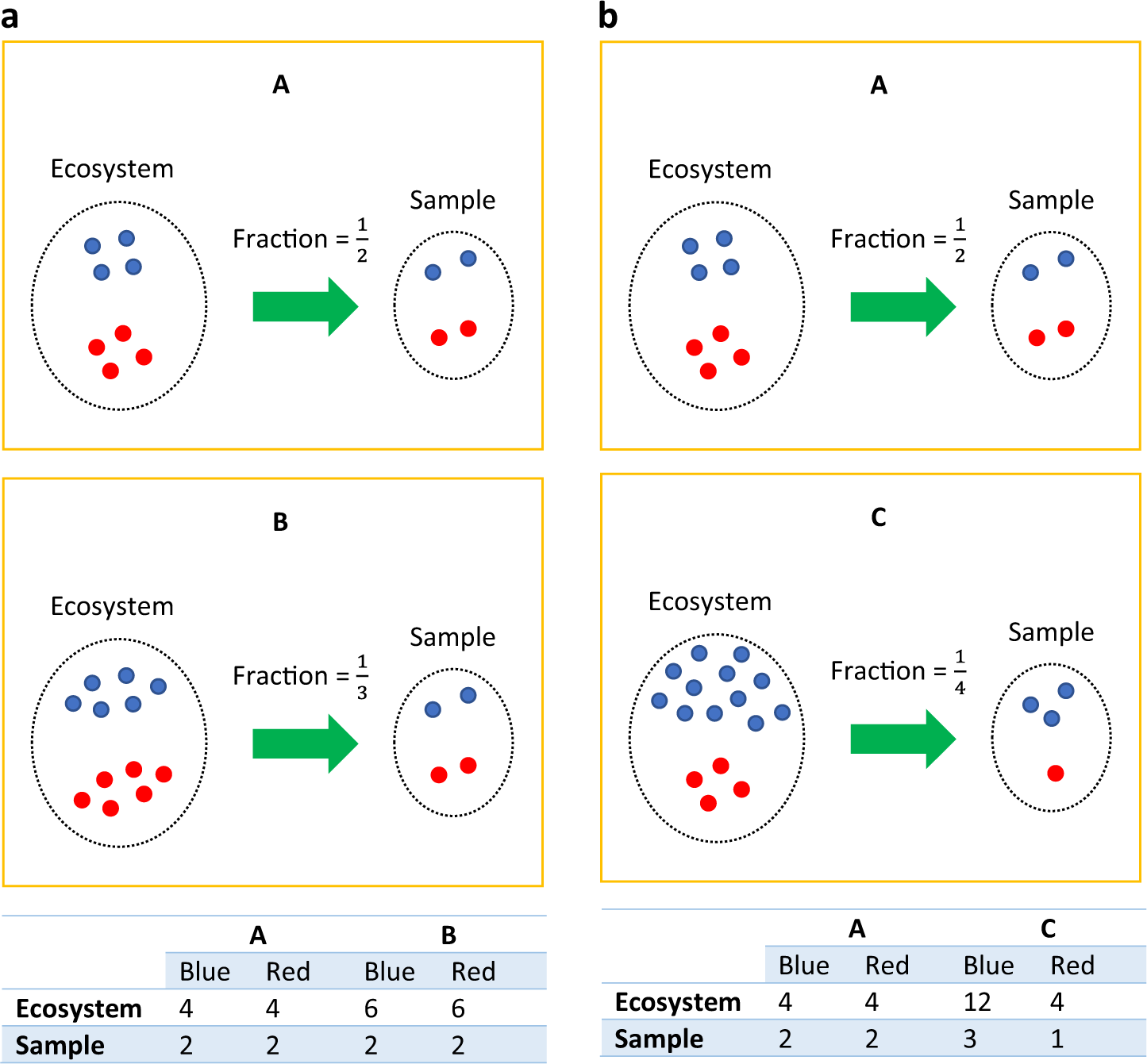
Analysis of microbial compositions: a review of normalization and differential abundance analysis | npj Biofilms and Microbiomes

The Statistical Analysis of Compositional Data” by John Aitchison (1986): A Bibliometric Overview - Carolina Navarro-Lopez, Salvador Linares-Mustaros, Carles Mulet-Forteza, 2022
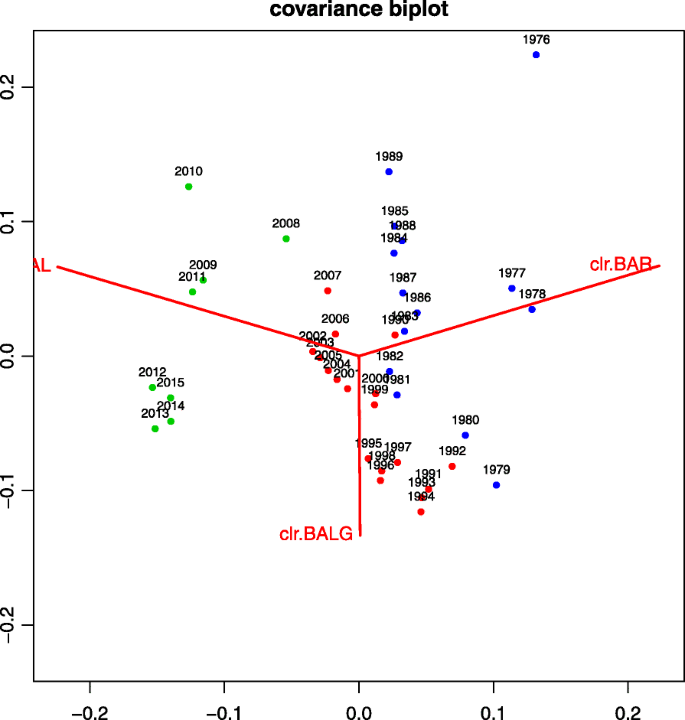
Compositional data techniques for the analysis of the container traffic share in a multi-port region | European Transport Research Review | Full Text
GitHub - belarius/matlab-compositional-analysis: Matlab functions for the statistical analysis of compositional data

Advances in Compositional Data Analysis: Festschrift in Honour of Vera Pawlowsky-Glahn | SpringerLink
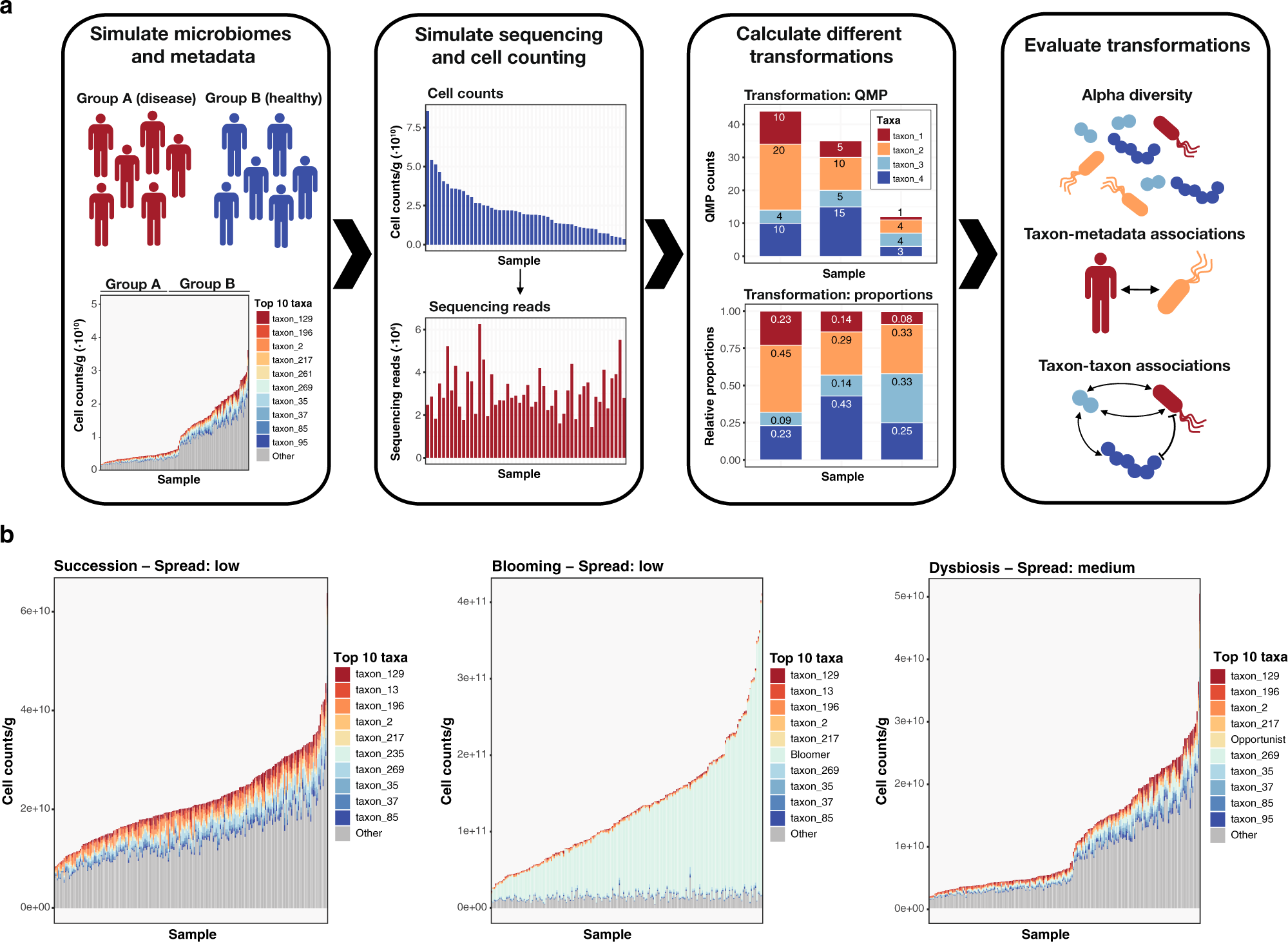
Benchmarking microbiome transformations favors experimental quantitative approaches to address compositionality and sampling depth biases | Nature Communications

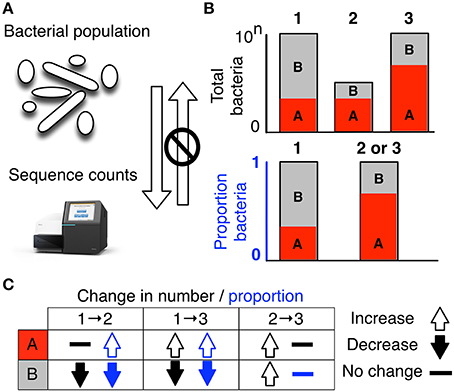












![PDF] Regression analysis with compositional data containing zero values | Semantic Scholar PDF] Regression analysis with compositional data containing zero values | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/1e4dc94d8b92acec92fb2537caa59c1b29a2c644/7-Figure1-1.png)